Have you ever tried to utilize python programming for Bio-Technology? When I realized there are a lot of opportunities available for Python, I got curious that whether can I use python for my domain Bio-Tech and I found there are plenty of opportunities available for Python programmers to automate their regular day-to-day tasks.
So I thought to share how did I start the basic codings to calculate the percentage of ATGC content and complimentary sequence to nucleotide sequence in Python?
Module creation:
The python modules are a collection of codes containing functions, variables, etc. These stored codes are highly reusable and can be imported and accessed whenever necessary.
- Read the nucleotide sequence and calculate the length:

Step 1.1: Create a function called ‘sequence length’ with the sequence as the input parameter (‘seq’).
Step 1.2: Then the total length of the sequence is calculated using the length function.
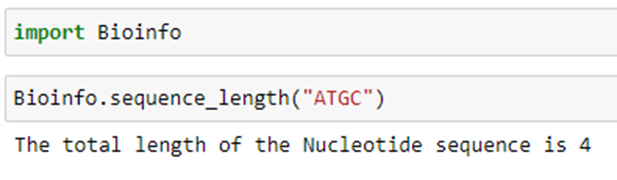
2. Calculated the percentage of A, T, G, and C content from the nucleotide sequence:

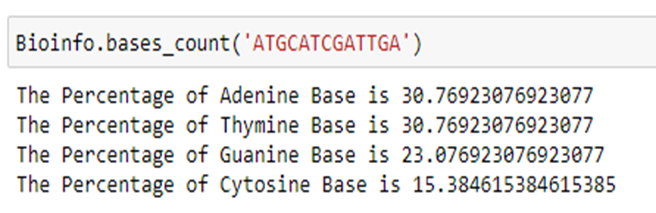
Step 2.1: I have created a function called ‘bases_count ’with the sequence as the input parameter (‘seq’).
Step 2.2: The input sequence is passed inside the for loop and it checks for each base. If the base is A, the ‘count_a’ is incremented to 1. Likewise, the remaining bases are checked, and the individual count variable will be updated.
Step 2.3: From the obtained individual base count, the percentage of each base from the input sequence can be calculated using ‘(‘count’ / ‘seq length’) * 100’. Sequence length is the total length of the sequence and count is the count of each base.
3. Complimentary sequence for the input sequence:
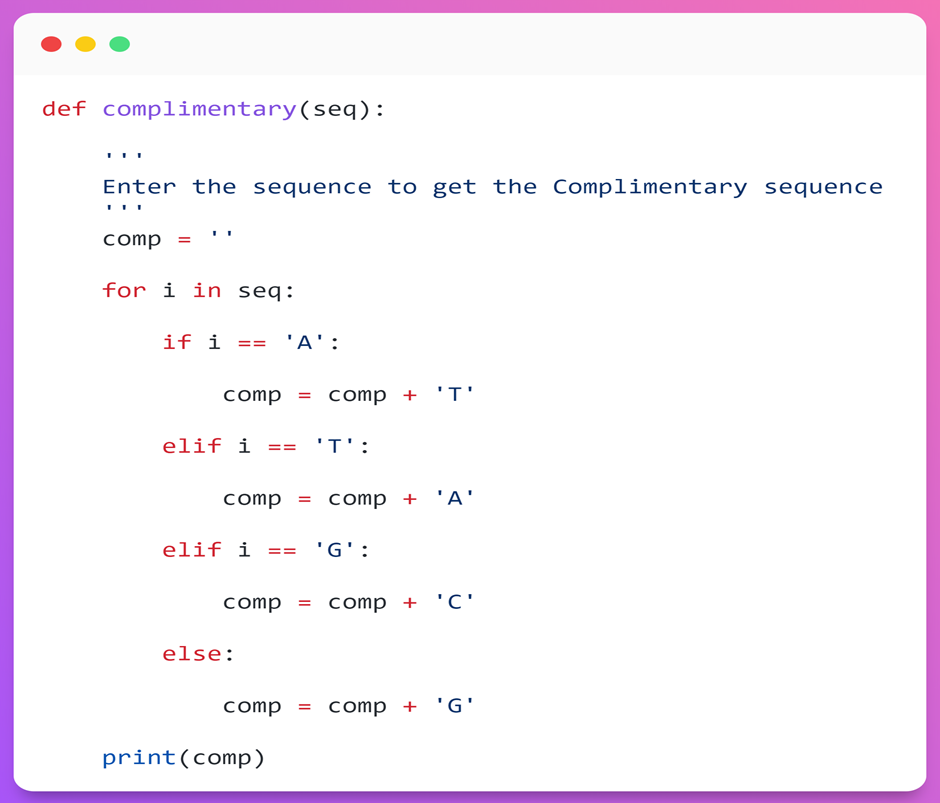

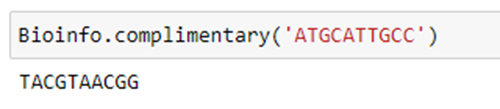
Step 3.1: I have created a function called complimentary with the sequence as the input parameter (‘seq’).
Step 3.2: The input is passed inside the for loop and it checks each base. If the base is ‘A’, it returns ‘T’ and is added to the ‘comp’ variable. Likewise, the remaining bases are checked, and the ‘comp’ variable will be updated.
Step 3.3: Then, it displays the sequence in the ‘comp’ variable, which is complimentary to the DNA sequence. That is (A: T and G: C).
That’s it! What you’re waiting for? Just start practicing the above python code to get get a percentage of ATGC content and complimentary sequence (i.e., A: T and G: C) for the input sequences
I strongly recommend you write the code to create simple tasks in Bio-Technology like this. It helps you to understand the programming concepts and then move to some high-level tasks in the future.
In case you’re looking for Data Science in the Bio-Tech domain follow me since I am already exploring a lot in that.
If you have any quires feel free to connect me on LinkedIn www.linkedin.com/in/aswindsc
Thanks & Regards
Aswin Balamurugan


